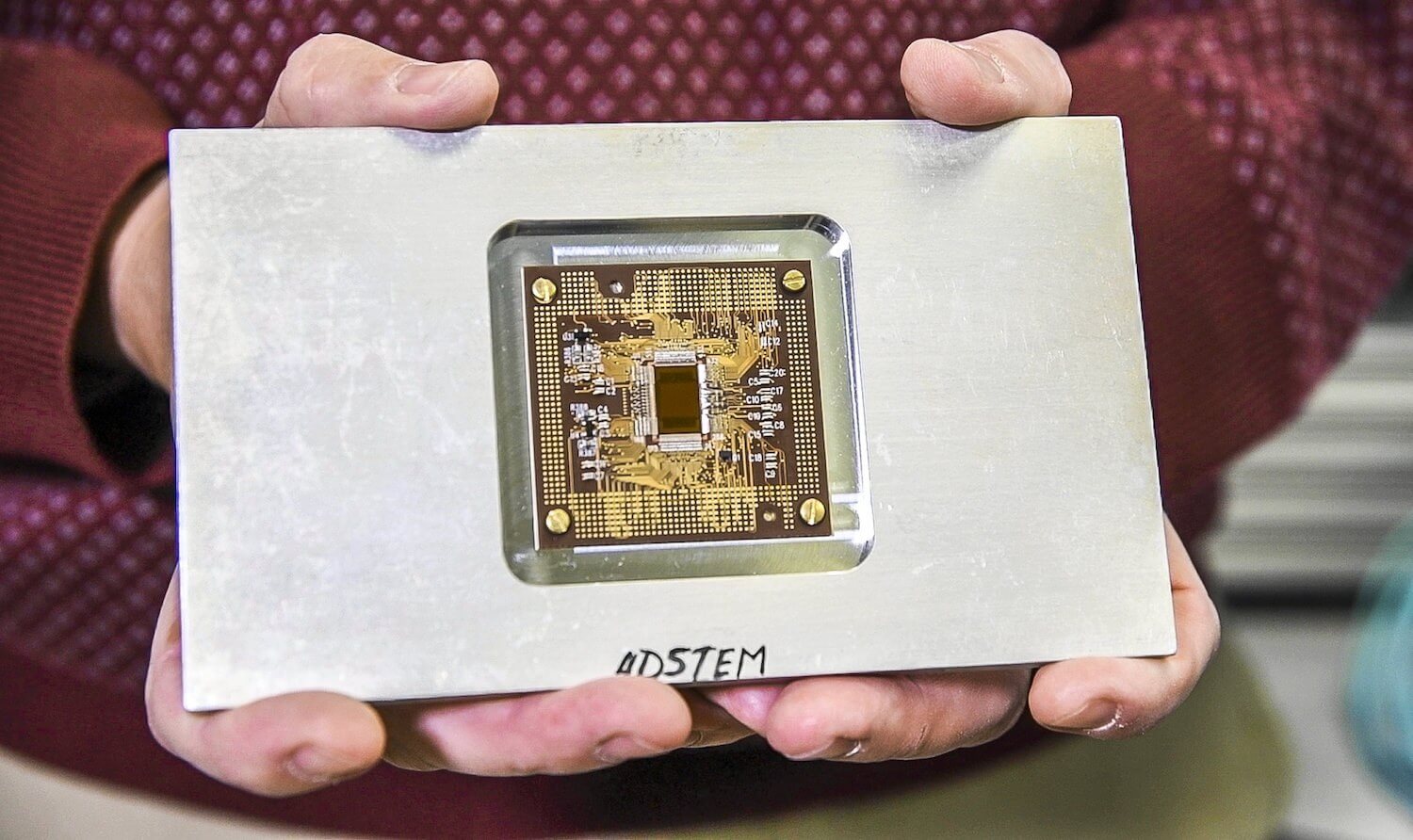
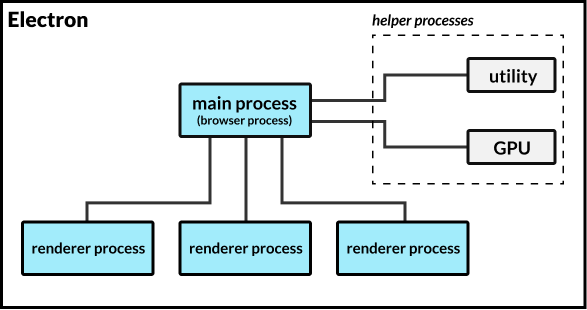
![PDF] GPU acceleration of all-electron electronic structure theory using localized numeric atom-centered basis functions | Semantic Scholar PDF] GPU acceleration of all-electron electronic structure theory using localized numeric atom-centered basis functions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c5fb18d1fd88a58b7724d77d404a257acc1f77a1/6-Figure1-1.png)
PDF] GPU acceleration of all-electron electronic structure theory using localized numeric atom-centered basis functions | Semantic Scholar

DOE CSGF 2022: Quantum Chemistry at Scale: Multi-Node Multi-GPU Two-Electron Integral Code Gener... - YouTube

Inq, a Modern GPU-Accelerated Computational Framework for (Time-Dependent) Density Functional Theory | Journal of Chemical Theory and Computation
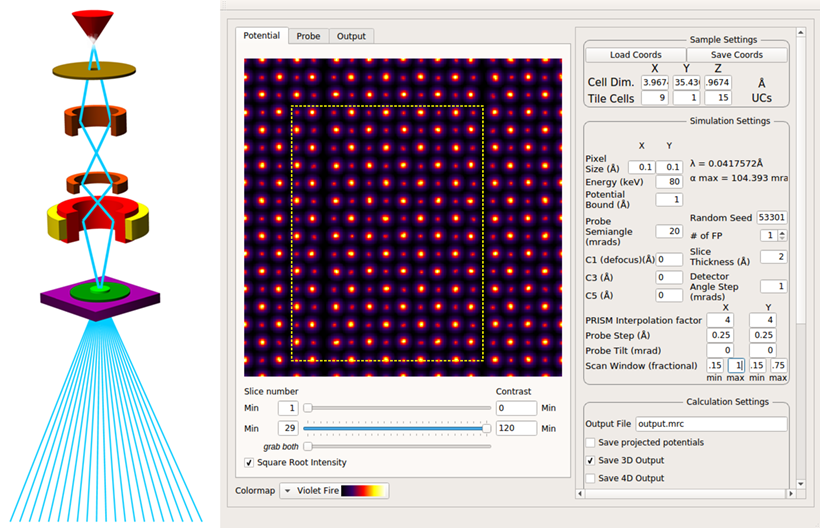
Prismatic Software For STEM Simulation - Documentation, tutorials, and source code for the software package Prismatic, a GPU implementation of image simulation algorithms in scanning transmission electron microscopy (STEM)

Ben Sandofsky on Twitter: "I loaded the same Slack channel in Safari, their Electron app, and a sideloaded copy of their iOS app on my M1 Mac. Electron: 481 MB Safari: 749

Theia Scientific using balena and Grafana for Machine Learning-Powered Real-Time Image Analysis with NVIDIA Jetson AGX Orin
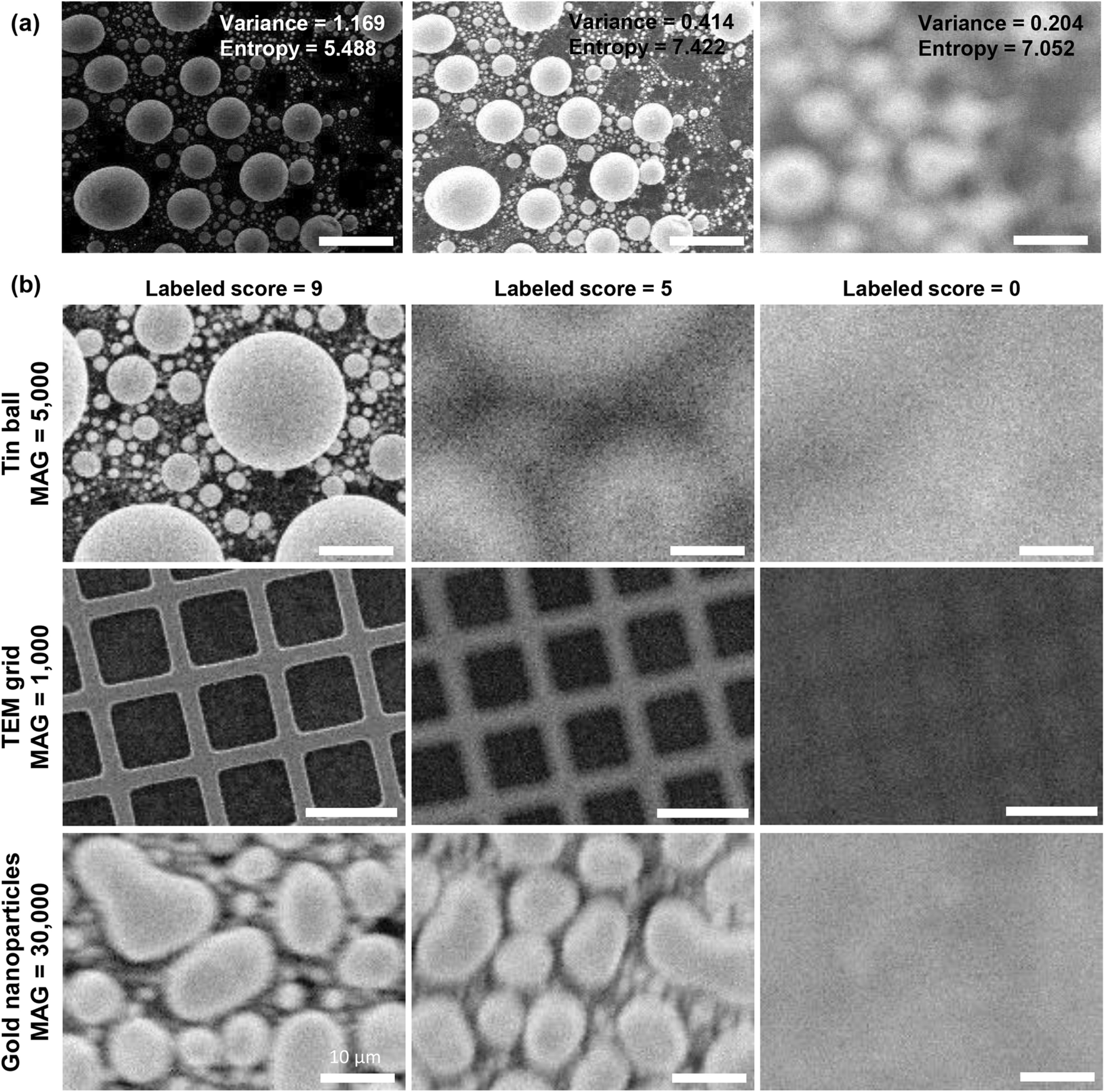

Robust autofocusing for scanning electron microscopy based on a dual deep learning network | Scientific Reports
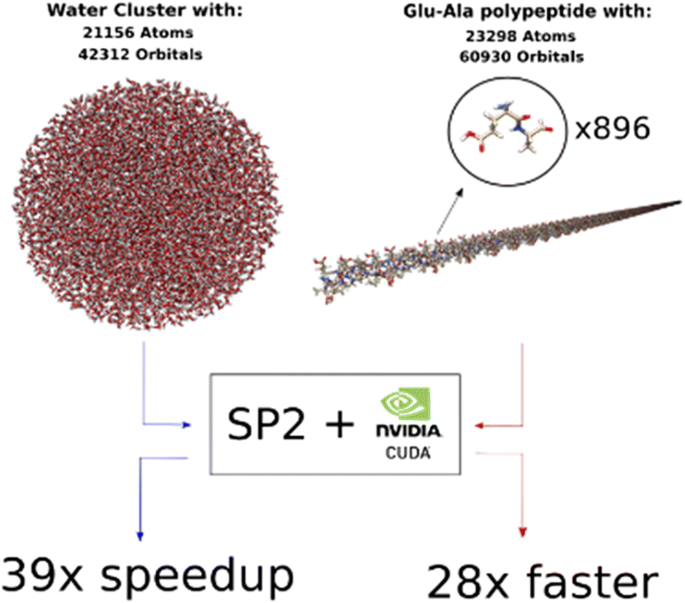
GPU algorithms for density matrix methods on MOPAC: linear scaling electronic structure calculations for large molecular systems | SpringerLink

CF1015H12D FD10015M12D RX 590 580 480 470 570 GPU Cooler Fan For Sapphire RX470 RX590 RX580 RX480 RX570 NITRO SpecialEdition Fan|Fans & Cooling| - AliExpress